Alkaline soil affecting crop production
Alkaline soil affecting crop production
Plants are a major source of iron for human nutrition. Unfortunately, iron is frequently a limiting nutrient for plant growth with low iron availability caused by high soil pH being one of the broadest ranging abiotic stresses in agriculture. Around 30% of arable land is too alkaline for optimal crop production. In order to solve the problem of iron deficiency, it is desirable to breed plants that have increased iron in those parts that are consumed by humans, and that are tolerant of high pH soil. To do this, we must first understand the molecular basis of iron uptake, transport, and storage in plants.
Ionomics
We use bionomics to explore the plant metal homeostasis mechanism. The ionome is defined as all the mineral nutrient and trace elements found in an organism, representing the inorganic component of cellular and organismal systems (Lahner et al., 2003). Ionomics is the study of the ionome using high-throughput elemental analysis technologies in combination with bioinformatic and genetic tools. Inductively coupled plasma spectroscopy (ICP-MS) was chosen for high throughput ion profiling due to its high sensitivity and large dynamic range, allowing simultaneous determination of all the mineral nutrients and trace elements in tissues. By profiling the mineral ion composition of the shoots and seeds of ecotypes and mutants of Arabidopsis thaliana, we have established an efficient way to uncover the network of the ionome and its interactions. An online searchable database called PiiMS (www.purdue.edu/dp/ionomics) for has been established (Baxter et al., 2007).
Two interstesting mutants
The analysis of fast neutron mutagenized plants has allowed the
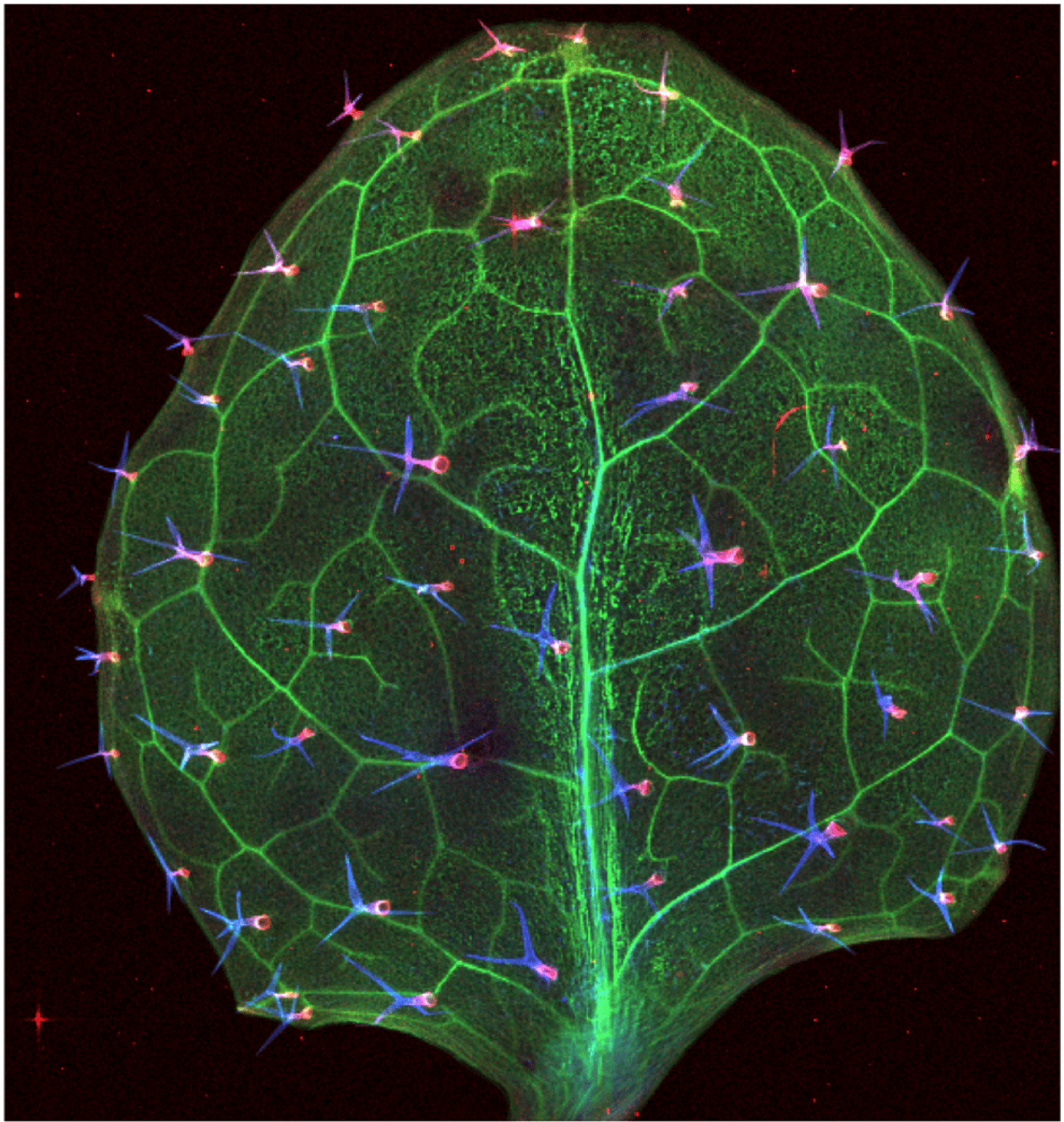
identification of 51 confirmed mutants (Lahner et al., 2003). Two fast neutron mutants, 131:61 and 118:65, with decreased and increaed iron accumulation in seeds respectively, were identified using ionomics to analyze shoots and seeds of ecotypes and mutants. 131:61 exhibits interveinal chlorosis, and produces seeds with lower Fe content than wild type when grown on regular soil. Moreover, 131:61 cannot set seeds when grown on alkaline soil . Despite the iron deficiency phenotype, 131:61 has the same concentration of iron in its shoots as wild type plants under both Fe sufficient and deficient conditions, and synchrotron x-ray fluorescence computed microtomography shows that 131:61 accumulates more iron in the vasculature and trichomes of leaves suggesting a defect in iron mobilization out of the vasculature rather than in iron uptake from soil. In contrast, 118:65 is able to thrive and has a higher iron concentration in its seeds than wild type when grown on alkaline soil. 118:65 exhibits strong rhizosphere acidification that may explain the ability to thrive on alkaline soil, although the increased acidification response does not appear to be due to an increase in transcript levels for any of the Fe regulated AHA genes. 118:65 shows tolerance to Fe deficiency as well as to high Zn, Ni, Co, and Cd stresses. Hopefully, the genes we identify will be useful for improving the iron content of crop seeds and/or tolerance to iron limitation.