 Project Leader:
Project Leader:
Mary Lou Guerinot, Ph.D.
Associate Director
Toxic Metals Superfund Research Program
Professor, Biological Sciences
Dartmouth College
Project Co-Leader:
David E. Salt, Ph.D.
Professor of Genome Enabled Biology, University of Nottingham Nottingham, UK
Rice, a staple food for over half the world’s population, represents a significant dietary source of arsenic, a known cause of cancer. It is vital that strategies to reduce arsenic in rice are developed, and establishing the way that arsenic reaches, and accumulates within, the edible parts of the rice grain is key to this endeavor.
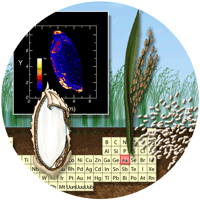
The long-term goal of this project is to prevent arsenic accumulation in the edible portion of the rice grain, but the work will also potentially provide information on genes responsible for transporting arsenic and other contaminant metals or metalloids into the tissues of other edible plant parts. The presence of essential or contaminant metals in living tissue is typically the result of transport proteins, which operate under tight genetic regulation. We will study how rice moves arsenic into the seeds as they develop by imaging elemental distribution under a range of exposures during grain development. We use elemental imaging techniques to map where the arsenic is within the plant, the grain, and even within individual cells, and X-ray spectroscopy to show its chemical form.
Recent studies show that both rice and Arabidopsis thaliana roots actively efflux intracellular arsenic back into the soil, yet the effluxer(s) responsible remains elusive. To identify the gene(s) responsible for this efflux, we utilized Fluorescently-Activated-Cell-Sorting (FACS) together with RNA-sequencing (RNA-seq) to generate genome-wide expression maps for epidermal, cortex, and endodermal cell types of A. thaliana roots exposed to arsenic.
In addition, we are screening 526 Multi-Parent Advanced Generation Integrated Cross (MAGIC) A. thaliana lines for variation in arsenic tolerance caused by the genomic variation in this population. Analysis of the variation has produced quantitative trait loci (QTL) responsible for arsenic tolerance. These QTL, together with the cell-type specific expression data, provide two distinct, high resolution data-sets capable of identifying not only genes responsible for arsenic efflux back to the soil, but other means of arsenic detoxification in plant roots.
Ultimately, we aim to understand the genetic control of arsenic homeostasis in plants so that we can develop plants that do not accumulate arsenic.
How does arsenic get into our food? Why does rice accumulate arsenic?
Project Background
Publications
Mary Lou Guerinot Pubmed
David Salt Pubmed